Figure 2
A Resurgence of the Idea of Hypertriglyceridemia and Lower Serum (HDL-C) as Predictive Factors for Insulin Resistance (IR) & Type 2 Diabetes Mellitus Development: A Narrative Review
Kulvinder Kochar Kaur*
Published: 24 July, 2025 | Volume 9 - Issue 1 | Pages: 001-012
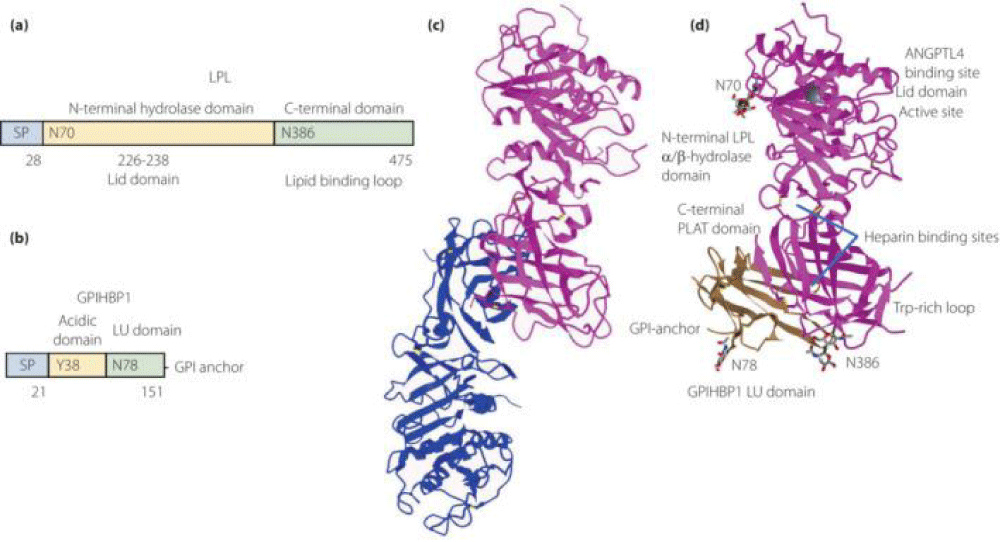
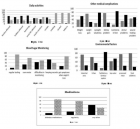
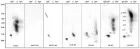
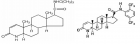
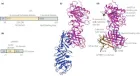
Figure 2:
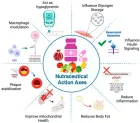
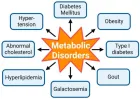
Courtesy ref no-51The structure of lipoprotein lipase (LPL) homodimer and LPL–LPL-LPL-LPL-LPL-glycosylphosphatidylinositol-anchored high‐density lipoprotein binding protein 1 (GPIHBP1) complex. (a) N‐terminal hydrolase with Lid domain and C‐terminal domain with lipid‐binding loop of LPL. Signal peptide (SP) and N‐glycosylation sites (N70 and N386) are shown. (b) Acidic domain and cysteine‐rich LU (Ly6/uPAR) domain, and C‐terminal hydrophobic regions of GPI‐anchored protein are shown. Y38 is modified by sulfation, and N‐glycosylation site (N78) is shown. (c) Ribbon model structures of Bos taurus LPL homodimer are obtained from PDB ID: 8ERL at MMDB (NCBI). Head‐to‐tail homodimer formation was thought to be critical for LPL secretion and the maintenance of the enzymatic activity, whereas recent data suggested that LPL homodimer is present in vitro, and LPL is synthesized and secreted as a monomer in vivo. (d) Ribbon model structures of human LPL/GPIHBP1 complex (pink and brown) are obtained from PDB ID: 6E7K at MMDB (NCBI). N‐terminal LPL α/β‐hydrolase domain and C‐terminal PLAT (polycystin‐1, lipoxygenase, α‐toxin) domain of LPL and GPIHBP1 LU domain are shown. Portions of angiopoietin‐like protein (ANGPTL) binding site, the lid domain covering the catalytic active site, heparin binding sites, and Trp‐rich loop in LPL are also shown. N‐terminal sequences (residues 21–61) containing the disordered acidic domain of GPIHBP1 are not defined in the ribbon diagram.
Read Full Article HTML DOI: 10.29328/journal.niogb.1001022 Cite this Article Read Full Article PDF
More Images
Similar Articles
-
Hypoglycemia by Ginseng in type 2 Diabetic Patient: Case ReportAngelo Michele Carella*,Teresa Marinelli,Armando Melfitano,Michele Di Pumpo,Matteo Conte,Angelo Benvenuto. Hypoglycemia by Ginseng in type 2 Diabetic Patient: Case Report . . 2017 doi: 10.29328/journal.hodms.1001001; 1: 001-006
-
Cardiac Autonomic Modulation in children and Preadolescents obeseMário Augusto Paschoal*. Cardiac Autonomic Modulation in children and Preadolescents obese. . 2017 doi: 10.29328/journal.hodms.1001002; 1: 007-009
-
The interaction between photonic technology and physical exercise: The action of low-level laser therapyAntonio Eduardo de Aquino Jr*,Fernanda Mansano Carbinatto. The interaction between photonic technology and physical exercise: The action of low-level laser therapy. . 2017 doi: 10.29328/journal.hodms.1001003; 1: 010-013
-
Behavioral factors of Abdominal Obesity and effects of lifestyle changes with Fiber AdequacyRoberto Carlos Burini*,Hugo Tadashi Kano,Mariana Santoro Nakagaki,Patricia Baston Frenhani,Katia Cristina Portero-McLellan. Behavioral factors of Abdominal Obesity and effects of lifestyle changes with Fiber Adequacy. . 2017 doi: 10.29328/journal.hodms.1001004; 1: 014-022
-
Study of body composition, physiological variables in Grade III obese submitted to arm ergometer testRenato Vidal Linhares*,Felipe Monnerat Marino Rosa,José Quaresma,João Regis Ivar Carneiro,Valéria Bender Braulio,Roberto Simão,Denise Pires de Carvalho,osé Fernandes Filho. Study of body composition, physiological variables in Grade III obese submitted to arm ergometer test . . 2017 doi: 10.29328/journal.hodms.1001005; 1: 023-029
-
Upper gut bacterial overgrowth is a potential mechanism for Glucose Malabsorption after Vertical Sleeve GastrectomyTimothy R Koch*,Timothy R Shope,Matthew E Sharbaugh. Upper gut bacterial overgrowth is a potential mechanism for Glucose Malabsorption after Vertical Sleeve Gastrectomy. . 2017 doi: 10.29328/journal.hodms.1001006; 1: 030-035
-
Body mass index in a group of security forces (policemen). Cross-sectional studyGuillermo Padrón Arredondo*. Body mass index in a group of security forces (policemen). Cross-sectional study. . 2018 doi: 10.29328/journal.niogb.1001007; 2: 001-004
-
Herbal approach for obesity managementPreeti Singh*. Herbal approach for obesity management. . 2018 doi: 10.29328/journal.niogb.1001008; 2: 005-016
-
Dietary and Lifestyles assessment among Obese Women in Gaza City, PalestineMarwan O Jalambo*,Basil Kanoa,Mohammed S Ellulu,Smaher Younis,Mueen El-Kariri. Dietary and Lifestyles assessment among Obese Women in Gaza City, Palestine. . 2018 doi: 10.29328/journal.niogb.1001009; 2: 017-025
-
Obesity-Treatment by drugsSahithi G*. Obesity-Treatment by drugs. . 2019 doi: 10.29328/journal.niogb.1001010; 3: 001-001
Recently Viewed
-
From Wallets to Waistlines: Exploring the Link between Family Income (Wage Distribution) and Obesity Indices in Pakistani PopulationsAtif Akbar,Waqas Ghulam Hussain*. From Wallets to Waistlines: Exploring the Link between Family Income (Wage Distribution) and Obesity Indices in Pakistani Populations. New Insights Obes Gene Beyond. 2026: doi: 10.29328/journal.niogb.1001024; 10: 001-014
-
Screening for Depressive Symptoms in Clinical and Nonclinical Youth: The Psychometric Properties of the Dutch Children’s Depression Inventory-2 (CDI-2)Denise HM Bodden*,Yvonne Stikkelbroek,Daan Creemers,Sanne PA Rasing,Elien De Caluwe,Caroline Braet. Screening for Depressive Symptoms in Clinical and Nonclinical Youth: The Psychometric Properties of the Dutch Children’s Depression Inventory-2 (CDI-2). Insights Depress Anxiety. 2025: doi: 10.29328/journal.ida.1001047; 9: 028-039
-
Knowledge, Attitude, and Practice of Healthcare Workers in Ekiti State, Nigeria on Prevention of Cervical CancerAde-Ojo Idowu Pius*, Okunola Temitope Omoladun, Olaogun Dominic Oluwole. Knowledge, Attitude, and Practice of Healthcare Workers in Ekiti State, Nigeria on Prevention of Cervical Cancer. Arch Cancer Sci Ther. 2024: doi: 10.29328/journal.acst.1001038; 8: 001-006
-
Oral Cancer Management is not just Treatment! But also, how early Pre-cancerous Lesions are Diagnosed & Treated!!Suresh Kishanrao*. Oral Cancer Management is not just Treatment! But also, how early Pre-cancerous Lesions are Diagnosed & Treated!!. Arch Cancer Sci Ther. 2024: doi: 10.29328/journal.acst.1001039; 8: 007-012
-
Breast Cancer in FemaleLorena Menditto*. Breast Cancer in Female. Arch Cancer Sci Ther. 2024: doi: 10.29328/journal.acst.1001040; 8: 013-018
Most Viewed
-
Physical Performance in the Overweight/Obesity Children Evaluation and RehabilitationCristina Popescu, Mircea-Sebastian Șerbănescu, Gigi Calin*, Magdalena Rodica Trăistaru. Physical Performance in the Overweight/Obesity Children Evaluation and Rehabilitation. Ann Clin Endocrinol Metabol. 2024 doi: 10.29328/journal.acem.1001030; 8: 004-012
-
Hypercalcaemic Crisis Associated with Hyperthyroidism: A Rare and Challenging PresentationKarthik Baburaj*, Priya Thottiyil Nair, Abeed Hussain, Vimal MV. Hypercalcaemic Crisis Associated with Hyperthyroidism: A Rare and Challenging Presentation. Ann Clin Endocrinol Metabol. 2024 doi: 10.29328/journal.acem.1001029; 8: 001-003
-
Effects of dietary supplementation on progression to type 2 diabetes in subjects with prediabetes: a single center randomized double-blind placebo-controlled trialSathit Niramitmahapanya*,Preeyapat Chattieng,Tiersidh Nasomphan,Korbtham Sathirakul. Effects of dietary supplementation on progression to type 2 diabetes in subjects with prediabetes: a single center randomized double-blind placebo-controlled trial. Ann Clin Endocrinol Metabol. 2023 doi: 10.29328/journal.acem.1001026; 7: 00-007
-
Exceptional cancer responders: A zone-to-goDaniel Gandia,Cecilia Suárez*. Exceptional cancer responders: A zone-to-go. Arch Cancer Sci Ther. 2023 doi: 10.29328/journal.acst.1001033; 7: 001-002
-
Ectopic adrenal tissue at the spermatic cordAbdallah Chaachou,Nizar Cherni,Wael Ferjaoui*,Mohamed Dridi,Samir Ghozzi. Ectopic adrenal tissue at the spermatic cord. J Clin Med Exp Images. 2022 doi: 10.29328/journal.jcmei.1001024; 6: 001-002

If you are already a member of our network and need to keep track of any developments regarding a question you have already submitted, click "take me to my Query."